Department of Physical and Life Sciences, Nevada State College, Henderson, NV USA
Department of Zoology, Ohio Wesleyan University, Delaware, OH USA
Biology Department, University of Detroit Mercy, Detroit, MI USA
ReBUILDetroit, University of Detroit Mercy, Detroit, MI USA
Abstract
Genetic screens provide a mechanism to identify genes involved with different cellular and organismal processes. Using a Flp/FRT screen in the Drosophila eye we identified mutations that result in alterations and de-regulation of cell growth and division. From this screen a group of undergraduate researchers part of the Fly-CURE consortium mapped and characterized a new allele of the gene Hippo, HpoN.1.2.
Description
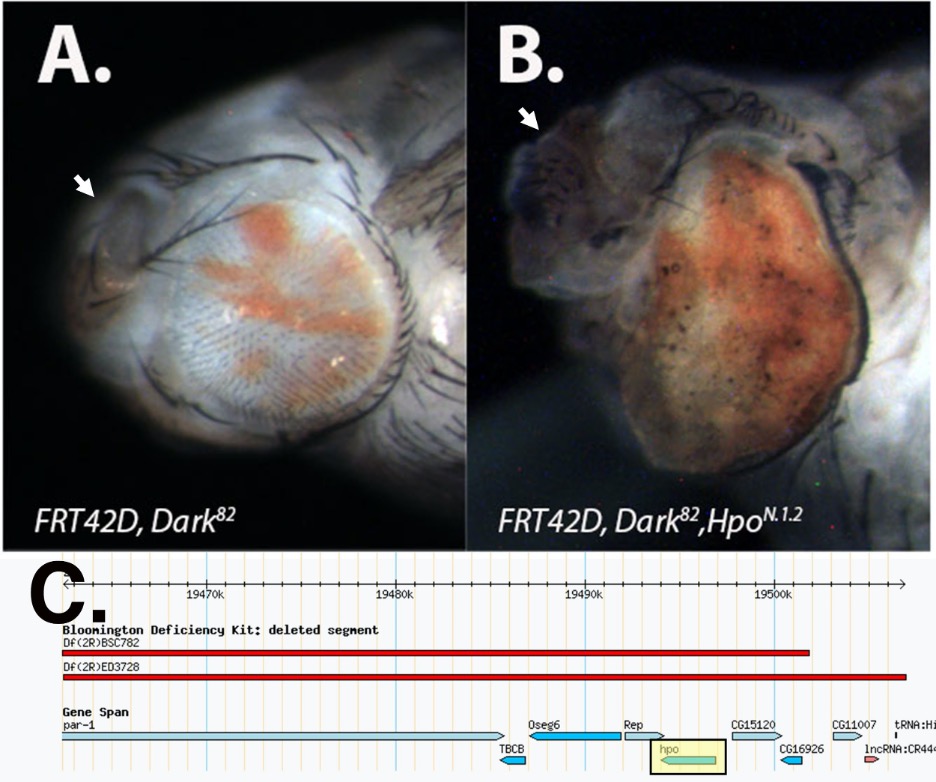
In order to identify conditional regulators of cell growth and tumorigenesis, an EMS-based genetic screen was conducted in Drosophila melanogaster utilizing the Flp/FRT system in an apoptotic null background (Akdemir et al., 2006). A fly line possessing the FRT42D, Dark82 chromosome and containing a mini-white cassette (w+mC) was used for EMS mutagenesis. Subsequent matings to FRT42D; Ey-Flp flies facilitated phenotypic screening in eye tissue. Phenotypes observed in the screen included alterations in mosaicism (red > white pigmentation pattern), eye and/or antennal overgrowth, defects in patterning, and pupal lethality (Kagey et al. 2012). One of the pupal lethal mutants identified in this screen, N.1.2, is discussed here. Genetic crosses between FRT42D, Dark82, N.1.2 X FRT42D; Ey-Flp resulted in near complete pupal lethality (~ 90%) due to dramatic tissue overgrowth of the eye, antennae, and inter-ocular space (Figure 1B compared to 1A). Due to the pupal lethality associated with the eye tissue overgrowth, flies were dissected from late pupal stages in order to visualize the phenotype. Imaging following dissection shows that the mutant eyes are comprised of mostly mutant (pigmented) tissue that protrudes from the eye cavity creating tissue folding (Figure 1B). Control FRT42D, Dark82 flies exhibited a balance in the red:white ratio and no eye overgrowth (Figure 1A). For direct comparison to N.1.2, the FRT42D, Dark82 mosaic eye was also imaged at the late pupal stage. Eyes were imaged under 70% ethanol on an AM Scope digital camera at 40x using a LED light ring.
In order to map the genetic location of the N.1.2 mutation, we conducted complementation tests with deficiency strains, and looked for lethality, consistent with the phenotype of homozygous mutant flies of the N.1.2 stock. Deficiency crosses were conducted by mating FRT42D, Dark82, N.1.2/CyO virgin females to males from each of 86 deficiency stocks with deletions distal to the FRT42D site on the right arm of chromosome two. All stocks used for mapping were part of the Bloomington Deficiency 2R Kit (Cook et. al., 2012). The genetic mapping was conducted by undergraduate research students from Nevada State College, Ohio Northern University, Ohio Wesleyan University, and the University of Detroit Mercy as part of the Fly-CURE consortium (Vrailas-Mortimer et al. 2021; Bieser et al. 2019; Stamm et al. 2019). The N.1.2 mutant failed to complement two of these deficiencies, Df(2R)BSC782 (2R: 19,451,027..19,501,804) and Df(2R)ED3728 (2R:19,462,450..19,726,747). These two deficiencies overlapped from cytological bands 56D10-56D14 (2R:19,462,450..19,506,861) and included the hippo (hpo) gene locus and nine other protein coding genes (2R:19,492,996..19,496,856) (Figure 1C). Subsequent crosses to individual lethal alleles of hpo: hpoMGH1and hpoKS240, failed to complement N.1.2, confirming that the N.1.2 mutant is a newly isolated hpo allele, hpoN.1.2 (Harvey et al., 2003; Udan et al., 2003).
The hpo gene functions as a negative regulator of cell growth and is a part of a highly conserved signaling pathway first characterized in Drosophila (reviewed in Harvey and Hariharan 2012). Overall, research has shown that the hippo pathway is critical for regulating organ size through regulation of apoptosis, cell survival, cell polarity, and cell proliferation (Harvey et.al., 2003; Jia et.al. 2003; Pantalacci et. al., 2003; Udan et. al., 2003; reviewed in Yu et. al, 2015). Consistent with the eye phenotypes observed in the N.1.2 mutation, mutations in hpo are known to result in striking overgrowth phenotypes in a variety of tissue types from flies to humans (Pan 2010, Pluoffe et al., 2015; Yu et. al., 2015). Identification of this novel hpoN.1.2 allele will support further research into the molecular mechanisms by which multicellularity is regulated and restricted by this critical signaling pathway.
Reagents
FRT42D, Dark82/CyO (Akdemir et al., 2006)
FRT42D, Dark82, hpoN.1.2/CyO (this manuscript)
FRT42D; Ey-Flp (BDSC 8211)
Bloomington Drosophila Stock Center 2R Deficiency Kit (Cook et al., 2012)
w1118; Df(2R)BSC782/SM6a (BDSC 27354)
w1118; Df(2R)ED3728/SM6a (BDSC 9067)
yw; FRT42D, hpoKS240/CyO (BDSC 25085)
hpoMGH1 (Harvey et al. 2003)
Acknowledgments
Stocks obtained from the Bloomington Drosophila Stock Center (NIH P40OD018537) were used in this study.
References
Funding
Fly-CURE (K. Bieser, J. Kagey) National Science Foundation IUSE Award (NSF 2021146). A. Murry is funded by the NIH ReBUILDetroit (TL4GM118983)
Reviewed By
AnonymousHistory
Received: February 20, 2021Revision received: March 29, 2021
Accepted: March 29, 2021
Published: April 8, 2021
Copyright
© 2021 by the authors. This is an open-access article distributed under the terms of the Creative Commons Attribution 4.0 International (CC BY 4.0) License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.Citation
Siders, JL; Bieser, KL; Hamill, DR; Acosta, EC; Alexander, OK; Ali, HI; Anderson, MJ; Arrasmith, HR; Azam, M; Beeman, NJ; Beydoun, H; Bishop, LJ; Blair, MD; Bletch, B; Bline, HR; Brown, JC; Burns, KM; Calagua, KC; Chafin, L; Christy, WA; Ciamacco, C; Cizauskas, H; Colwell, CM; Courtright, AR; Diaz Alavez, L; Ecret, RI; Edriss, F; Ellerbrock, TG; Ellis, MM; Extine, EM; Feldman, E; Fickenworth, LJ; Goeller, CM; Grogg, AS; Hernandez, Y; Hershner, A; Jauss, MM; Jimenez Garcia, L; Franks, KE; Kazubski, ET; Landis, ER; Langub, J; Lassek, TN; Le, TC; Lee, JM; Levine, DP; Lightfoot, PJ; Love, N; Maalhagh-Fard, A; Maguire, C; McGinnis, BE; Mehta, BV; Melendrez, V; Mena, ZE; Mendell, S; Montiel-Garcia, P; Murry, AS; Newland, RA; Nobles, RM; Patel, N; Patil, Y; Pfister, CL; Ramage, V; Ray, MR; Rodrigues, J; Rodriquez, VC; Romero, Y; Scott, AM; Shaba, N; Sieg, S; Silva, K; Singh, S; Spargo, AJ; Spitnale, SJ; Sweeden, N; Tague, L; Tavernini, BM; Tran, K; Tungol, L; Vestal, KA; Wetherbee, A; Wright, KM; Yeager, AT; Zahid, R; Kagey, JD (2021). Genetic Mapping of a new Hippo allele, HpoN.1.2, in Drosophila melanogaster. microPublication Biology. 10.17912/micropub.biology.000383.Download: RIS BibTeX