Department of Chemistry, Birmingham-Southern College, Birmingham, AL 35254, USA
Abstract
Low solubility in aqueous solutions is a significant limitation of the otherwise promising anticancer ruthenium complex KP1019. In laboratory studies, this challenge is often overcome by using DMSO to help drive the drug into solution. Since DMSO was previously shown to alter the bioactivity of platinum-based chemotherapeutics, here we examine DMSO’s effects on KP1019. Using Saccharomyces cerevisiae as a model organism, we apply multiple measures of growth inhibition to demonstrate that DMSO reduces the drug’s toxicity. This reduction in bioactivity correlates with spectrophotometric changes consistent with DMSO-dependent increases in the stability of the KP1019 pro-drug. The impact of DMSO on the biology and chemistry of KP1019 suggests this solvent should not be used in studies of this and similar anticancer ruthenium complexes.
Description
In an early stage dose-escalation clinical trial, anticancer ruthenium complex indazolium trans-[tetrachlorobis(1H-indazole)ruthenate(III)] (also known as KP1019), stabilized disease progression in five of six evaluable patients (Hartinger et al. 2008). However, KP1019’s low solubility prevented identification of the maximum tolerated dose. In laboratory studies, the challenge of KP1019’s poor solubility is often overcome by dissolving the drug in dimethyl sulfoxide (DMSO) before diluting it into the growth medium or buffer being used in a particular assay (Bergamo et al. 2009; Singh et al. 2014; Golla et al. 2017; Śpiewak et al. 2019). Though many studies control for the biological impacts of DMSO itself by including a vehicle only control, the impacts of DMSO on KP1019’s chemistry and bioactivity have not been directly tested. Notably, a previous study showed that the activities of the platinum drugs cisplatin and carboplatin are reduced by 87 to 98% respectively when solubilized with DMSO (Hall et al. 2014). Given this finding and ruthenium’s high affinity for sulfur containing ligands, we hypothesized that DMSO would also affect KP1019.
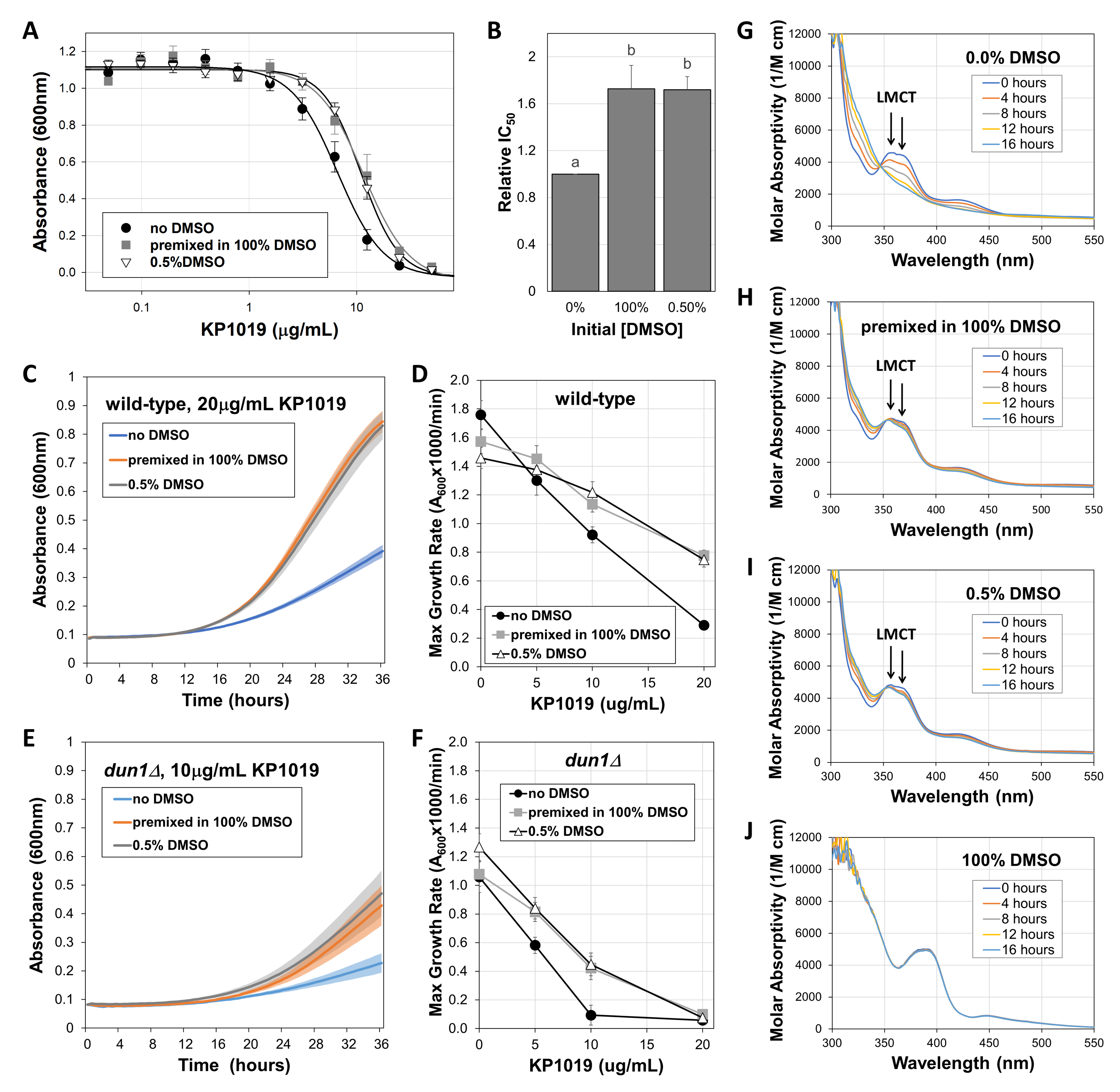
To determine whether DMSO increases or decreases KP1019 bioactivity, dose-response growth inhibition assays were conducted with wild-type Saccharomyces cerevisiae, which is an appropriate model for studying KP1019, because both budding yeast and cancer cells experience similar effects from the drug. For example, KP1019 damages DNA and induces oxidative stress in both cancer and yeast cells (Kapitza et al. 2005; Golla et al. 2017; Stultz et al. 2020). Additionally, multidrug resistant cancer and yeast cell lines both remain sensitive to KP1019 treatment (Heffeter et al. 2005; Stevens et al. 2013). Since prior studies dissolved KP1019 in DMSO before dilution in aqueous solutions, we compared this approach to dissolving KP1019 directly in yeast media. As seen in Figure 1A, premixing KP1019 with pure DMSO shifts the dose response curve to the right, significantly increasing growth at multiple KP1019 concentrations. Consistent with a recent study, which reported the IC50 for KP1019 in the absence of DMSO as 6.8µg/mL (Stultz et al. 2020), here we found a similar value of 6.7±0.8µg/mL (mean ± S.E., n=6) for the same, widely used wild-type strain BY4742. More importantly, initially dissolving the drug in DMSO prior to dilution in yeast media significantly increased the IC50 to 11.4±1.8µg/mL.
To determine whether this loss of efficacy was dependent on initially dissolving KP1019 in pure DMSO, we also conducted the growth inhibition assay with drug that was dissolved directly in media containing a final concentration of 0.5% DMSO. This approach also reduced KP1019’s efficacy, raising the IC50 to 11.2±0.9µg/mL, which is statistically comparable to the value obtained by first dissolving the drug in pure DMSO (p>0.05). These changes in efficacy are summarized in Figure 1B in which the IC50’s for each condition are normalized to the no DMSO control for each trial. These data illustrate that inclusion of DMSO increases KP1019’s IC50 by greater than 50%, representing a significant loss of bioactivity. This reduction in effective dose has potential to be clinically significant, as a roughly one-third drop in dose or dose intensity for other chemotherapies has been shown to worsen patient outcomes (Bonadonna et al. 1995; Budman et al. 1998)
As a complement to the end-point readings used for IC50 analyses, growth curve assays were conducted in the presence of select concentrations of KP1019 solubilized with or without DMSO. As seen in Figure 1C, when the drug was dissolved in 100% DMSO prior to dilution in yeast media, or when KP1019 is dissolved in media containing 0.5% DMSO, yeast growth is substantially less inhibited with the average maximum growth rate increasing more than 2.5-fold from 0.29 ± 0.02 for the no DMSO control to 0.78 ± 0.04 and 0.75 ± 0.05 A600x1000/min (mean ± S.E., n=6), respectively. These results verify that DMSO reduces the toxicity of KP1019 in yeast. Likewise, other KP1019 concentrations resulted in dose-dependent decreases in the maximum growth rate of wild-type yeast (Figure 1D), and multivariate regression analyses verified that this growth inhibition was reduced in the presence of DMSO (p<0.05). Notably, there was not a statistically significant difference between the two approaches for including DMSO in the assay (p>0.05).
Analogous to observations in wild-type yeast, DMSO also reduced the efficacy of KP1019 against yeast lacking the kinase Dun1 (Figures 1E and 1F), which is essential for the yeast DNA damage response (Zhou and Elledge 1993). Whereas Figures 1D and 1F verify that dun1∆ yeast have a modest growth defect relative to wild-type strains even in the absence of drugs (Marek et al. 2013), Figures 1C-F verify previous observations that dun1∆ strains are hypersensitive to KP1019 (Singh et al. 2014; Stultz et al. 2020). For example, at 10µg/mL KP1019 in the absence of DMSO, the maximum growth rate of wild-type yeast is 0.92 ± 0.06 A600x1000/min (mean ± S.E.) (Figure 1D). Under these same conditions, the dun1∆ strain grows at roughly one tenth the rate of the wild-type, reaching a maximum of 0.09 ± 0.07 A600x1000/min (Figure 1F). In the dun1∆ mutant, the effects of DMSO are clear. For example, at 10µg/mL KP1019, inclusion of DMSO resulted in a more than four-fold increase in average growth rate, reaching 0.42 ± 0.08 and 0.45 ± 0.08 A600x1000/min when pre-mixed with 100% DMSO or dissolved in media containing 0.5% DMSO, respectively. However, these growth rates are still much lower than the wild-type controls. This sustained KP1019 sensitivity of the dun1∆ yeast suggests that even DMSO-impaired KP1019 can damage DNA. However, future studies will be required to establish whether DMSO interferes with KP1019’s other cellular impacts, such as proteotoxicity and oxidative stress (Kapitza et al. 2005; Stultz et al. 2020).
To determine whether DMSO’s effects on KP1019 bioactivity correlate with changes to the drug’s chemical properties, UV-vis spectra were analyzed. As seen in Figure 1G, the spectrum for KP1019 dissolved in buffer alone initially shows characteristic peaks at 357 nm and 421 nm, which are indicative of ligand to metal charge transfer (LMCT) (Kratz et al. 1994). These peaks gradually decrease over time as the drug’s chloride ligands are exchanged for water (Küng et al. 2001). When initially dissolved in DMSO then diluted in buffer (Figure 1H) or when dissolved in buffer already containing 0.5% DMSO (Figure 1I), the LMCT bands are largely maintained over the 16-hour timeframe of the experiment. As a control, we also tested KP1019 in pure DMSO for the duration of the experiment. As seen in Figure 1J, this condition shifts the LMCT peaks to the right (390 nm and 448 nm), and the spectrum does not change over the timeframe of the experiment. Altogether, these data may indicate that DMSO increases the stability of KP1019 in solution. This increased stability suggests that DMSO may interfere with KP1019’s activation by reduction, which is required for the metal to bind and damage biomolecules like DNA (Antonarakis and Emadi 2010). Though our data do not conclusively rule out formation of a highly stable complex that contains DMSO, the similarity of the initial spectra for aqueous solutions with and without DMSO (Figures 1G-I), make ruthenium binding directly to DMSO an unlikely explanation in this case, as such binding would be anticipated to alter the LMCT bands. Instead DMSO may have a strong solvent interaction with KP1019, which could prevent the reduction required for activation and binding to cellular components (Antonarakis and Emadi 2010).
In light of KP1019’s limited aqueous solubility, the sodium salt analog KP1339 was developed (Trondl et al. 2014). This drug contains the same active metal complex but has a sodium counterion rather than indazolium, making KP1339 far more soluble than KP1019. Even still, some studies of KP1339 have used DMSO as a vehicle (Flocke et al. 2016; Bakewell et al.. 2020), and we predict this solvent would have similar effects on KP1339 as we report here for KP1019. Given DMSO’s impact on KP1019, caution should be used when comparing studies that do and do not dissolve the drug in DMSO, and when possible, future research on ruthenium complexes should avoid using DMSO as a vehicle.
Methods
Request a detailed protocolDrug Synthesis
KP1019 (CHEBI ID: 77760) was synthesized as described previously (Stevens et al. 2013).
Determination of IC50 and Growth Curve Analysis
KP1019’s ability to inhibit yeast growth was quantified in terms of an IC50 as described previously with the following modifications (Stevens et al. 2013). KP1019 was dissolved in SDC yeast media, dissolved in 100% DMSO and then diluted in SDC with a final DMSO concentration of 0.5%, or dissolved in SDC containing 0.5% DMSO. To maintain a consistent vehicle concentration, drug solutions were two-fold serially diluted in media with or without 0.5% DMSO. Overnight Saccharomyces cerevisiae cultures were normalized to A600 0.1, then diluted 20-fold more in SDC with or without 0.5% DMSO as appropriate. An equal volume of cell suspension was added to the serially diluted KP1019. Relative concentrations of KP1019 were estimated with a pre-incubation scan at 360 nm, and trials where different experimental groups (DMSO vs. no DMSO) were more than 10% away from the mean were excluded from further analyses. Plates were incubated at 30°C for 20 hours before end-point absorbance (600nm) was measured. A similar approach was used for growth curve assays, except mid-log phase cultures were used to reduce the duration of lag phase, microtitre plate lids were pre-treated to prevent accumulation of condensation (Brewster 2003), plates were incubated at room temperature, and absorbance was measured at 30-minute intervals for 36 hours. Maximum growth rates were determined automatically by Biotek Gen5 software, which calculated the maximum slope by evaluating 5 points post lag phase.
UV-vis spectrophotometry
Absorbance spectra of solutions containing a final concentration of 0.25mM KP1019 were studied under four different conditions: dissolved directly in 0.5 M sodium phosphate buffer (pH 7.2) without DMSO, pre-dissolved in 100% DMSO then diluted to 0.5% DMSO in phosphate buffer, dissolved directly in phosphate buffer containing 0.5% DMSO, and dissolved in 100% DMSO only. Data was collected at 25°C every hour for 19 hours, and absorbance was converted to molar absorptivity by dividing by the concentration of KP1019. For clarity, representative scans at 0, 4, 8, 12, and 16 hours were graphed.
Reagents
The yeast strains used in this study were BY4742 (MATα his3Δ1 leu2Δ0 lys2Δ0 ura3Δ0) (Brachmann et al.. 1998) and a commercially available, isogenic dun1::kanMX strain (Winzeler et al. 1999).
References
Funding
Funding for this research provided by Furman University and Birmingham-Southern College.
Reviewed By
Anonymous and Oliver KerscherHistory
Received: May 19, 2021Revision received: July 29, 2021
Accepted: July 29, 2021
Published: August 4, 2021
Copyright
© 2021 by the authors. This is an open-access article distributed under the terms of the Creative Commons Attribution 4.0 International (CC BY 4.0) License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.Citation
Davis, J; Cetto, A; Campbell, M; Scoggins, S; Stultz, L; Hanson, P (2021). DMSO reduces the cytotoxicity of anticancer ruthenium complex KP1019 in yeast. microPublication Biology. 10.17912/micropub.biology.000436.Download: RIS BibTeX