Abstract
In nature, Drosophila melanogaster larvae are infected by parasitoid wasps and mount a cellular immune response to this infection. Several conserved signaling pathways have been implicated in coordinating this response, however our understanding of the integration and regulation of these pathways is incomplete. Members of the S1A serine protease family have been previously linked to immune functions, and our findings suggest roles for two S1A family members, CG10764 and CG4793 in the cellular immune response to parasitoid infection.
Description
Cellular immune responses are an important aspect of innate host defense against infection and are broadly conserved from insects to mammals. The model organism Drosophila melanogaster uses the cellular encapsulation response to protect against macroparasite infection (Carton et al., 2008; Mortimer, 2013). This response shows genetic conservation with human immune responses (Howell et al., 2012), and may serve as a useful model to better understand human immune cell functions. Drosophila larvae are commonly infected by parasitoid wasps and following infection mount a cellular immune response to kill the parasite. This response is mediated by two cell types, circulating macrophage-like immune cells known as plasmatocytes and infection-induced immune cells called lamellocytes (Honti et al., 2014; Rizki, 1957). Plasmatocytes operate as the first line responders to infection by recognizing and binding to the wasp egg (Mortimer et al., 2012; Russo et al., 1996). This process is then followed by the production of lamellocytes that form a consolidated multi-layered capsule, thereby killing the wasp (Kim-Jo et al., 2019; Russo et al., 1996). Recent findings have begun to elucidate the regulation of the encapsulation response in Drosophila, including a role for the evolutionarily conserved JAK-STAT signaling pathway (Sorrentino et al. 2004; Yang et al. 2015). The roles of JAK-STAT signaling are not completely understood, but the pathway has been linked to the production of lamellocytes (Bausek and Zeidler, 2014; Hanratty and Dearolf, 1993; Luo et al., 1995, 1997; Sorrentino et al., 2004).
Members of the S1A protease family are involved in many physiological processes, including the regulation of invertebrate immune responses (Cao and Jiang, 2018). In Drosophila, the S1A family is composed of more than 200 genes and includes the catalytically active serine proteases (SPs) and the serine protease homologs (SPHs), a group of SP-like proteins that are enzymatically inactive (Cao and Jiang, 2018). Many S1A family members have been linked to the antimicrobial immune response including the SP genes spirit, grass, psh and SPE, and the SPH genes sphe, sphinx1 and sphinx2 (Buchon et al., 2009; El Chamy et al., 2008; Kambris et al., 2006; Ligoxygakis et al., 2002; Patrnogic and Leclerc, 2017). However, the role of SP and SPH genes in regulating the fly antiparasitoid immune response is still not well-defined.
A recent study of transcriptional targets of JAK-STAT pathway activity showed that the S1A family members, the SP gene CG10764 (also known as SP77) and the SPH gene CG4793 (also known as cSPH128) are JAK-STAT pathway target genes (Bina et al., 2010). The JAK-STAT pathway is important for the production of lamellocytes following parasitoid infection (Sorrentino et al., 2004; Yang et al., 2015), and ectopic pathway activity leads to tumorigenesis as characterized by the precocious accumulation of lamellocytes (Ekas et al., 2010; Harrison et al., 1995). RNA interference (RNAi) mediated knock down of CG10764 and CG4793 in the JAK-STAT tumor model suggested that these genes may play antagonistic roles in regulating JAK-STAT signaling and lamellocyte production (Bina et al., 2010).
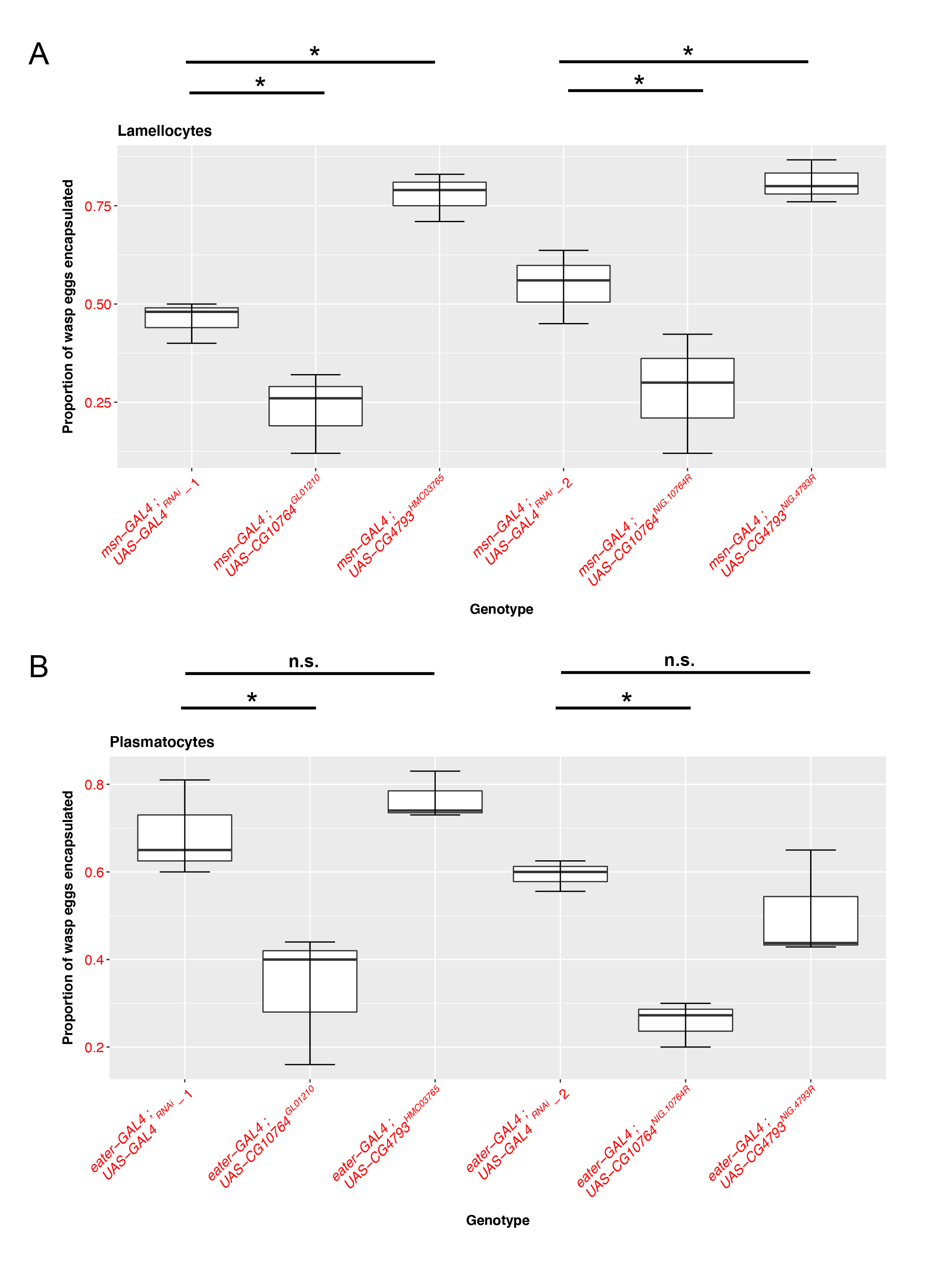
To evaluate the functional roles of these JAK-STAT regulated S1A family members in fly cellular immunity, we used two different RNAi lines with unique sequence targets to knock down each gene in both the plasmatocyte (using eater-GAL4) (Tokusumi et al. 2009a) and lamellocyte (using msn-GAL4) (Lam, et al. 2010; Tokusumi et al. 2009b) immune cell types and compared their ability to encapsulate parasitoid wasp eggs following infection. We find that knocking down CG10764 with either of the RNAi lines in lamellocytes (Figure 1A; UAS-CG10764GL01210: p= 0.00222, nEXP = 73, nCTRL = 72; UAS-CG10764NIG.10764R: p=0.0112, nEXP = 71 , nCTRL = 67) or plasmatocytes (Figure 1B; UAS-CG10764GL01210: p= 0.000955, nEXP = 75, nCTRL = 81 ; UAS-CG10764NIG.10764R: p= 1.74e-06, nEXP = 57 , nCTRL = 62) results in a significant reduction in the proportion of wasp eggs that are successfully encapsulated. These findings suggest that CG10764 may act as a positive regulator of encapsulation in both fly immune cell types. Conversely, RNAi-mediated knock down of CG4793 in lamellocytes with either of the RNAi lines results in a significant increase in encapsulation rate (Figure 1A; UAS-CG4793HMC03765: p= 1.09e- 05, nEXP, = 80, nCTRL = 72 ; UAS- CG4793NIG.4793R : p= 0.0098, nEXP, = 60, nCTRL = 67), but has no effect when knocked down in plasmatocytes (Figure 1B; UAS-CG4793HMC03765 : p= 0.56975, nEXP,=57, nCTRL =62 ; UAS- CG4793NIG.4793R : p= 0.321 , nEXP, = 57 , nCTRL = 62). This suggests that CG4793 may act as a negative regulator of encapsulation specifically in lamellocytes, the immune cell subtype that is induced following infection.
Based on our observations, we hypothesize that CG10764 and CG4793 play important and distinct roles in balancing immune activation. CG10764 appears to regulate the initiation of pro-immune signaling which triggers the host immune response against parasitoid infection. CG10764 likely encodes an active serine protease, and may influence immune activation through the direct cleavage of target proteins. On the other hand, CG4793 appears to be responsible for limiting the immune response when the defense mechanism is elicited. This is an important role which allows the host to avoid self-directed immune damage due to an overreactive immune system. CG4793 is an SPH gene and encodes a protein that is predicted to be catalytically inactive. However, these SPH proteins play regulatory roles in a variety of processes (Cao and Jiang, 2018), and it is likely that CG4793 is acting through a similar mechanism to limit immune activity. Thus, these S1A family members likely have cell-specific roles and regulate the cellular encapsulation process through distinct mechanisms.
A role for CG10764 and CG4793 in modulating JAK-STAT pathway activity has been previously demonstrated (Bina et al., 2010). Interestingly, these S1A family members were also shown to have opposing effects on the phenotype seen in hopTum flies, which display a melanotic phenotype due to ectopic JAK-STAT signaling (Bina et al., 2010; Hanratty and Dearolf, 1993; Luo et al., 1995). Here we show that CG10764 and CG4793 may also act antagonistically to maintain a balanced immune response and based on these previous studies, we hypothesize that this could potentially be via regulation of JAK-STAT signaling. However, a detailed mechanistic understanding of how these S1A family genes regulate cellular immunity and how their activity may be linked to JAK-STAT pathway signaling remain to be established. Additionally, further research into the human homologs of CG10764 and CG4793 may reveal conserved functions in human immunity and JAK-STAT mediated disease.
Methods
Request a detailed protocolDrosophila genetics. Tissue-specific modulation of gene expression can be achieved in D. melanogaster using the yeast-derived UAS-GAL4 system. GAL4 is a transcription factor that binds to the UAS enhancer sequence present in the promoter region controlling expression of the gene of interest (Brand and Perrimon, 1993). We used hemocyte specific GAL4 lines and two UAS-RNAi lines with distinct target sequences to knock down the genes of interest in each hemocyte type. UAS-GAL4RNAi was used as the control genotype. Independent control experiments were run with each UAS-RNAi experiment; UAS-GAL4RNAi-1 refers to the control replicates for experiments with the UAS-CG4793HMC03765 and UAS-CG10764GL01210 constructs and UAS-GAL4RNAi-2 refers to the control replicates for experiments with the UAS-CG10764NIG.10764R and UAS- CG4793NIG.4793R constructs. All Drosophila crosses were maintained on standard Drosophila medium (Molasses Formulation, Genesee Scientific) at 25C° on a 12 hour light:dark cycle.
Parasitoid wasp infection. For each genotype tested, approximately 25 virgin female GAL4 flies were mated with 10 UAS-RNAi line males. These crosses were transferred to egg lay chambers containing grape-juice plates (Genesee Scientific) supplemented with yeast paste and allowed to lay for 72 hours. For infection experiments, 25 F1 second instar larvae were picked from the egg lay plates and transferred into small petri dishes with standard Drosophila medium (Molasses Formulation, Genesee Scientific) together with 3 female LcNet wasps. All of the surviving larvae (~25/infection plate) were dissected 72 hours post infection and the number of encapsulated wasp eggs and live wasp larvae were counted. Each genotype for each experiment was performed in triplicate. All experimental crosses and infections were carried out at 25°C.
Encapsulation rate. After a 72 hour wasp exposure, larvae from each plate were dissected and scored for the presence of an encapsulated wasp egg or live wasp larva, to assay the encapsulation rate.
Data analysis and statistics. To analyze the effect of knockdown of proteases on wasp egg encapsulation rate, we used generalized linear models with quasibinomial errors to test for an effect of genotype, and then we performed Dunnett’s post hoc tests to compare each of the experimental genotypes to the control genotype. All statistics were done in the R statistical computing environment (R Core Team, 2020) using the “multcomp” (Hothorn et al., 2008), “plyr” package (Wickham, 2011). Graphs were produced using the “ggplot2” package (Wickham, 2009).
Reagents
The following Drosophila melanogaster stocks were used in this experiment:
UAS-RNAi lines:
| Short Genotype | Full Genotype | Stock ID |
| UAS-CG4793HMC03765 | y[1] v[1]; P{y[+t7.7] v[+t1.8]=TRiP.GL01210}attP40 | BDSC:41628 |
| UAS-CG10764GL01210 | y[1] sc[*] v[1] sev[21]; P{y[+t7.7] v[+t1.8]= TRiP. HMC03765}attP40 | BDSC:41628 |
| UAS-GAL4RNAi | y[1] sc[*] v[1] sev[21]; P{y[+t7.7] v[+t1.8]=VALIUM20-GAL4.1}attP2 | BDSC:35784 |
| UAS-CG10764NIG.10764R | P{NIG.10764R} | NIG:10764R-1 |
| UAS- CG4793NIG.4793R | P{NIG.4793R} | NIG:4793R-1 |
GAL4 lines (provided by Robert Schulz, University of Notre Dame):
| Genotype (FlyBase ID) | Expression | Reference |
| eater217-GAL4 (FBtp0057112) | Plasmatocytes | (Tokusumi et al. 2009a) |
| msn-GAL4 (FBtp0083721) | Lamellocytes | (Lam et al. 2010; Tokusumi et al. 2009b) |
We additionally used the figitid parasitoid wasp species Leptopilina clavipes (strain LcNet) (Mortimer et al. 2012) reared in the lab on Drosophila virilis.
Acknowledgments
We would like to thank members of the Mortimer lab for discussion of the project, and Robert Schulz and Tsuyoshi Tokusumi for providing Drosophila stocks. Stocks obtained from the Bloomington Drosophila Stock Center (Bloomington, IN, USA) (NIH P40OD018537) and the National Institute of Genetics Fly Stock Center (Mishima, Japan) were used in this study.
References
Funding
Research reported in this publication was supported by the National Institute Of General Medical Sciences of the National Institutes of Health under Award Number R35GM133760 to NTM, and a Grant-in-Aid of Research from the National Academy of Sciences administered by Sigma Xi, The Scientific Research Society to PKR.
Reviewed By
AnonymousHistory
Received: August 26, 2020Revision received: January 14, 2021
Accepted: February 11, 2021
Published: February 22, 2021
Copyright
© 2021 by the authors. This is an open-access article distributed under the terms of the Creative Commons Attribution 4.0 International (CC BY 4.0) License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.Citation
KR, P; Lee, J; Mortimer, NT (2021). The S1A protease family members CG10764 and CG4793 regulate cellular immunity in Drosophila. microPublication Biology. 10.17912/micropub.biology.000370.Download: RIS BibTeX